Questions about how to set up tau separation and tau scan.
Which is better, for instance. In our first trials, they seems to do the same thing but with different interfaces, the tau separation being more user friendly but the tauscan with the ability to add more channels.
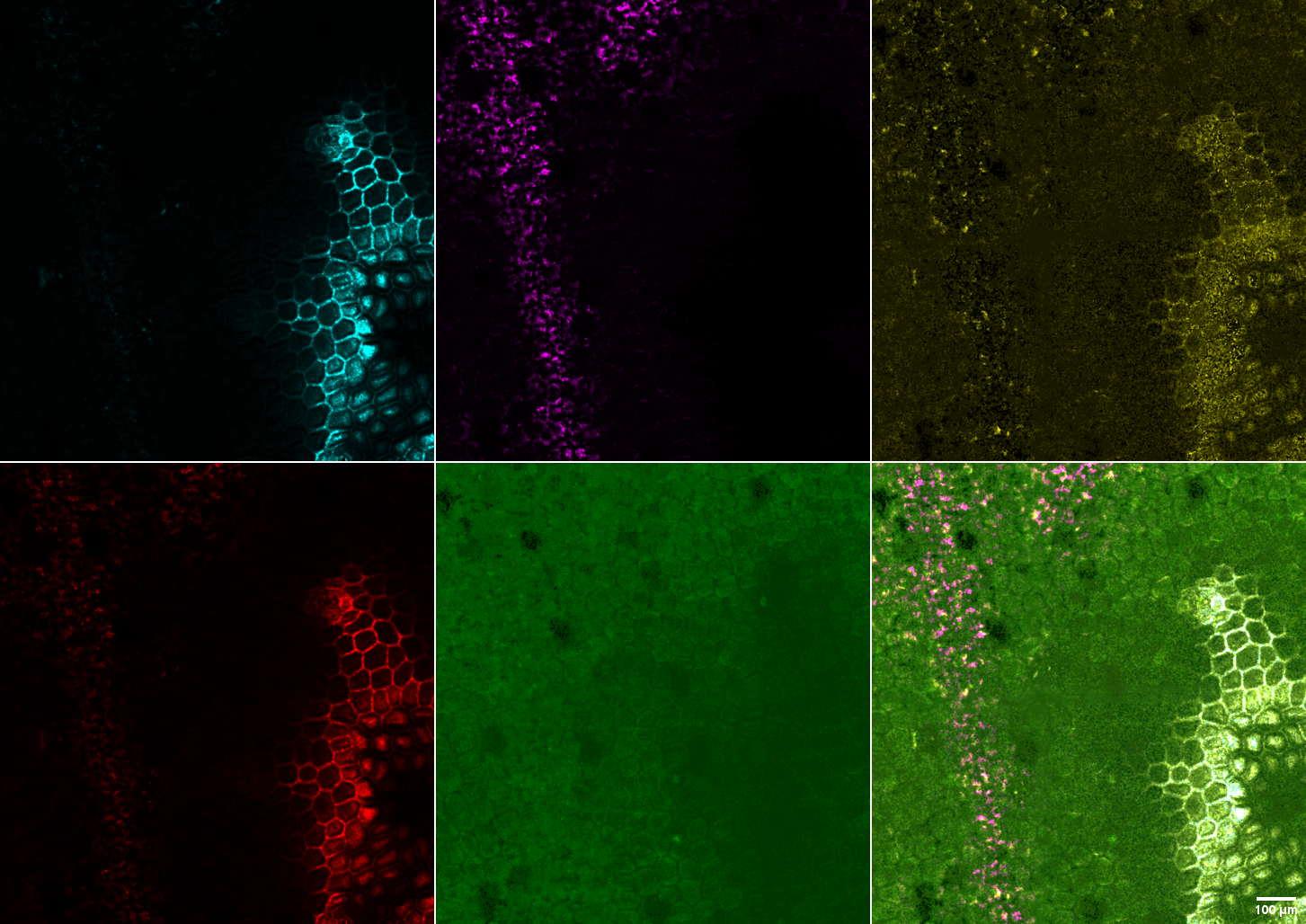
Test samples used here were pieces of plants. A leaf from a plant in the common area and a petal from a flower out in the rotary. Excitation was at approximately 550 nm and the detector was set with emission spectra up to approx 700 nm to collect most emitted photons of any energy level.

One question is which should we use, TauScan or TauSeparation? The two following histograms of arrival times appear essentially the same and the ability to move bins to collect the photons of a specific arrival time appear similar or the same.
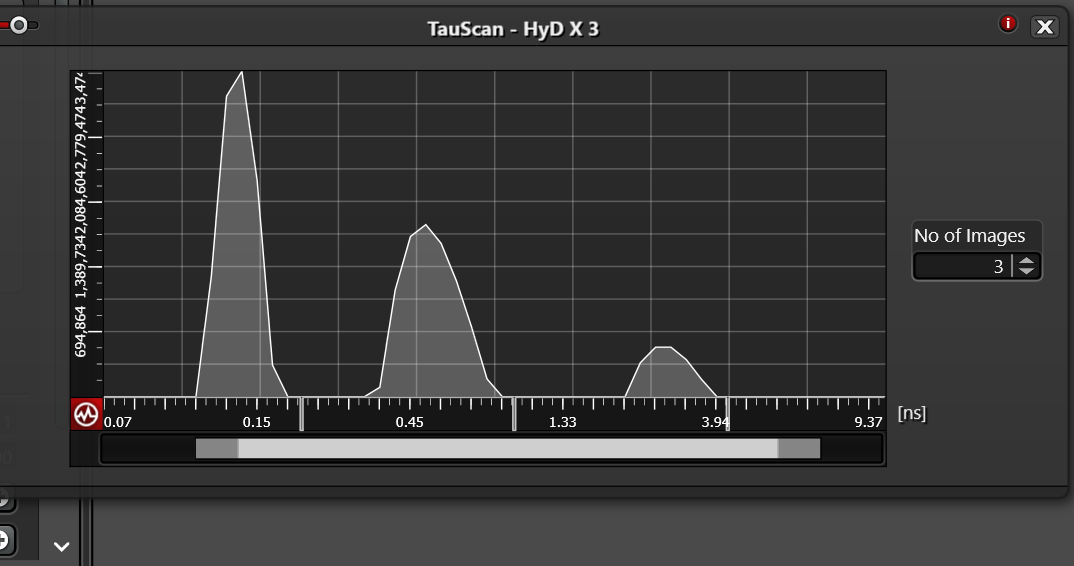
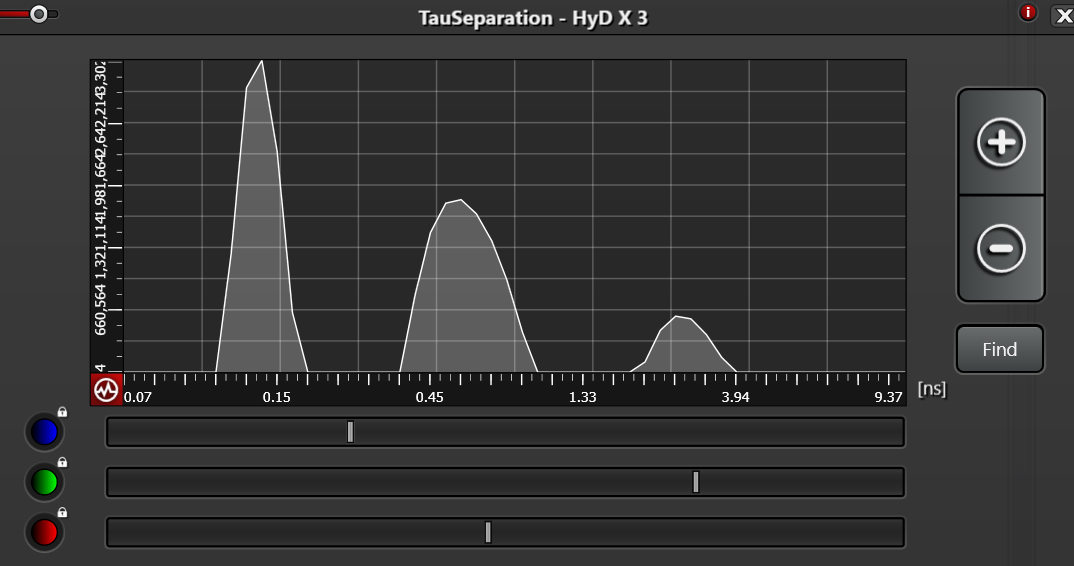
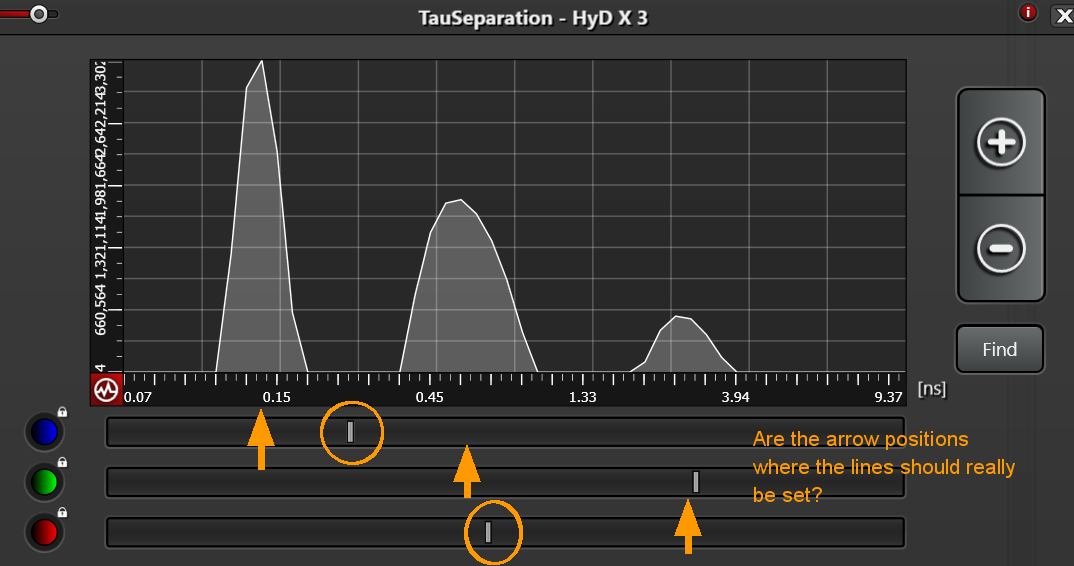
A big question is where to position the lines under the graph in TauSeparation. I would guess each should be directly below the peak of each curve. Is this correct?
In TauScan there appears to be no flexibility; the bins are of fixed width. Perhaps this answers the question which one to use (depending on the application).
Components were clearly separated.
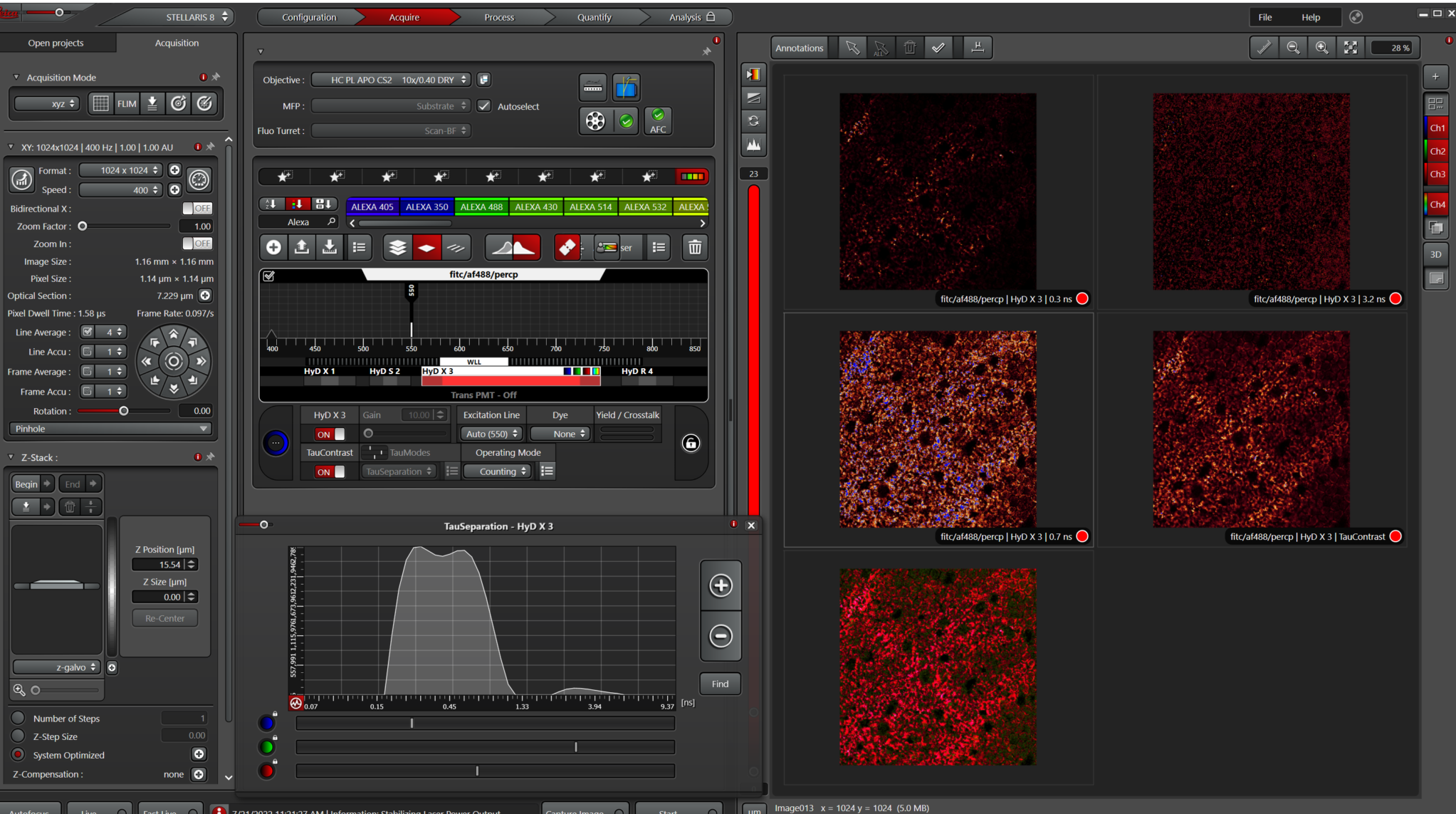
This example looks like the lines under the graph should be positioned at the peak of each component.
Unfortunately, sing BioFormats v6.5, one of the datasets would not open, but one of them did. We tried to decipher the metadata to find the lifetime bin widths for each image, but couldn't figure out which they were. These numbers do not appear to match the graphs.
ATLConfocalSettingDefinition|DetectorList|Detector|ImageChannel|DigitalGating|Begin 1.3272728E-09
ATLConfocalSettingDefinition|DetectorList|Detector|ImageChannel|DigitalGating|Duration 1.00848488E-08
ATLConfocalSettingDefinition|DetectorList|Detector|ImageChannel|DigitalGating|End 1.14121216E-08ATLConfocalSettingDefinition|DetectorList|Detector|TauChannel|TauChannelEntry|TauChannelValue 2.98022710767014E-09
Therefore, we do not know the lifetime bins for the next image. Each single color panel is a different lifetime. The cyan and red look very similar, but they are supposed to be differnt lifetimes. 10X N.A. 0.40 lens.